Chapter 4 Reproducibility
An important skill in data science is reproducibility. If you want to share your analysis you should describe the software you’ve used in a way that other people can easily install the same software as you have. There are two ways to easily do this, software environments and containerization. In software environments, software is installed in isolated folders and in containerization, software is installed in a virtual machine. Both of these methods work on linux.
4.1 Pixi based environments
If you want to consistently install software on a system without admin rights, pixi can help to determine 1) what programmes can be installed, 2) what programs you need to also install to get everying working, 3) manage these rules between operating systems and different versions.
Pixi is installed as:
# Install the binary
$ curl -fsSL https://pixi.sh/install.sh | bash
# Reset the shell
$ source ~/.bashrc
# Confirm installation.
$ pixi --version The documentation for pixi is found on https://pixi.sh/latest/basic_usage/.
Because putting all programs in a single database would get too massive, the developers put the software into groups, called channels. These channels can be general, such as conda-forge or domain specific, such as bioconda. Most bioinformatic software is found in the bioconda channel.
4.1.1 Usecase: installing qiime
The example I will give here with pixi is installing qiime, a software for analysing amplicon sequencing experiments. To find if this software is availible, we go to https://prefix.dev/. Then we type qiime in the search as so:
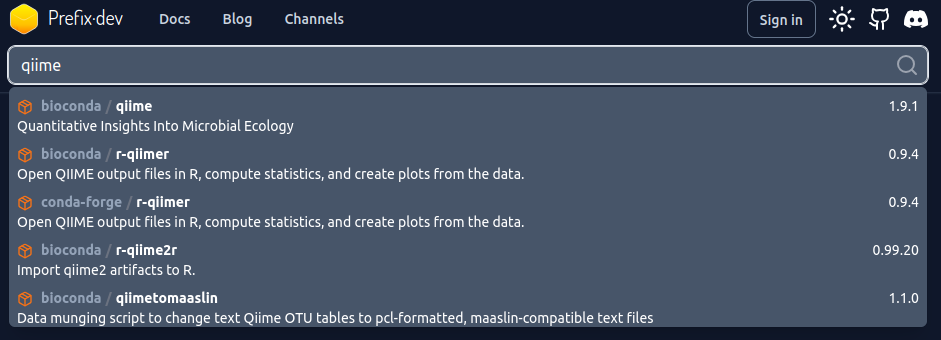
prefix.dev.Click on the first hit then you’ll see this:
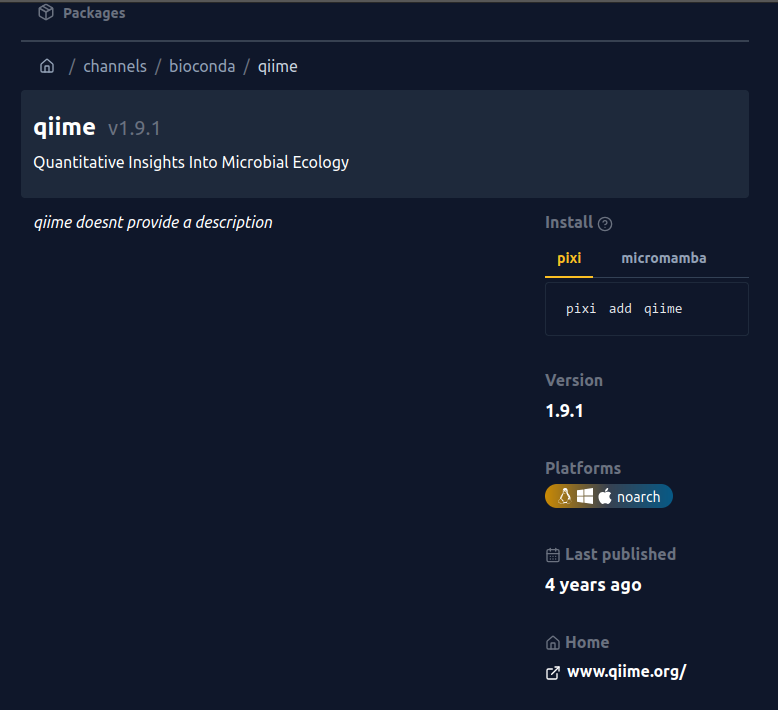
qiime software shows the version.Here you see that v1.9.1 is available. This means that you can install qiime!
Pixi has two modes of installation, global and local. The global installation places the program in the home (~/.pixi) directory. This makes the installed program behave as a program such as ls or echo. In the local method, you install the program for a specific pixi project. This is most useful when you mix programs with scripts and you want to share them. For instance, if you write a python program that you want to share with someone, you need to give the person the python code, and the python interpreter that you’ve used.
4.1.2 Global installation
For global installation, you would use the following command:
Here the -c bioconda indicates that qiime is located in the bioconda database.
4.1.3 Local installation
If you are writing an analysis using qiime and you want to share your analysis scripts written with bash, you’ll need to specify which qiime you used. You would do this as follows.
# Navigate to where you want your analysis
$ mkdir qiime-analysis
$ cd qiime-analysis
$ pixi init
✔ Created ~/qiime-analysis/pixi.tomlThen to specify that you want pixi to look in the bioconda database for the programs you type:
After this you can type
to install the latest version of qiime. Now you can use this programme (in qiime-analysis) by typing pixi run qiime.